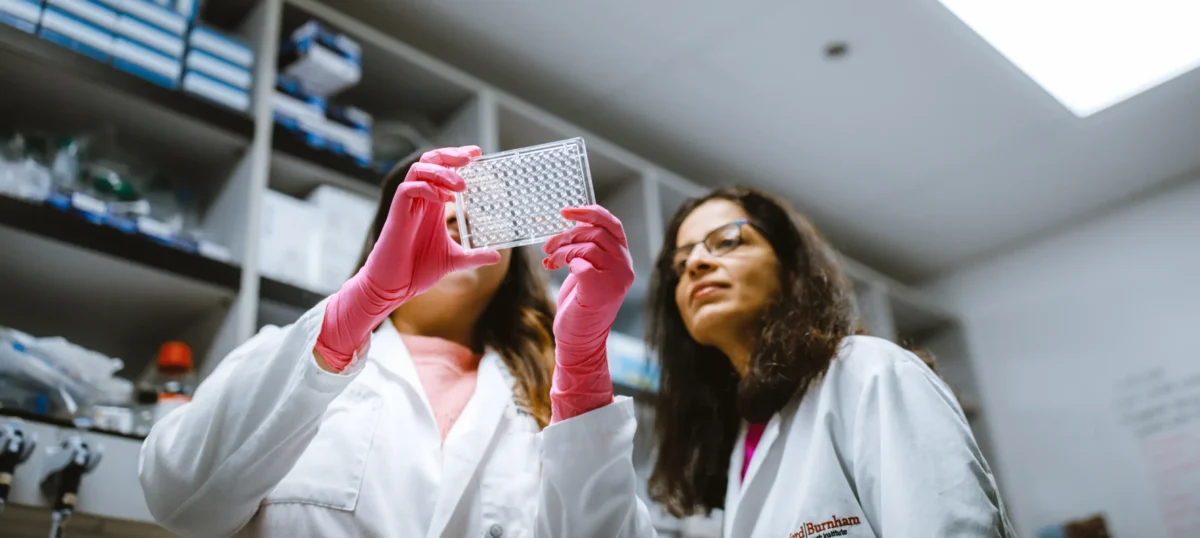
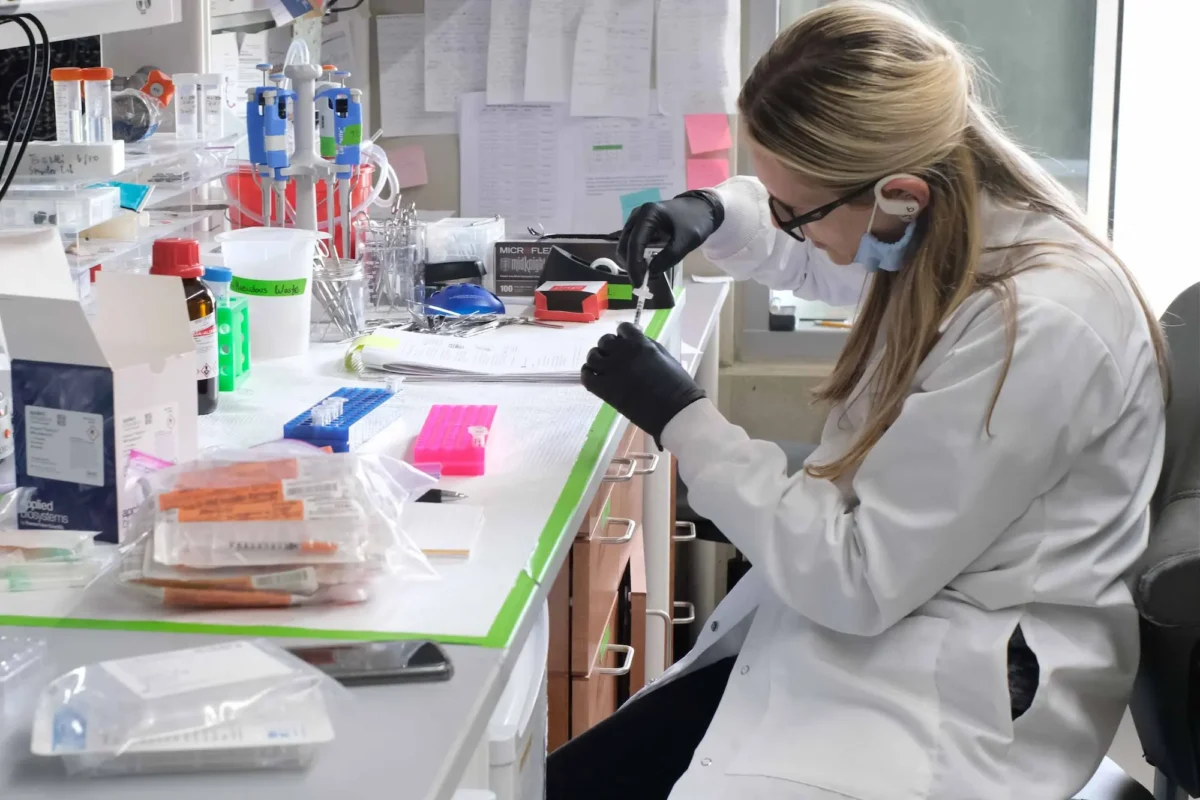
Disease-Focused Centers
Our centers combine research expertise and clinical collaboration to accelerate discovery, advance drug development and improve human health.
Enabling Technology Centers
State-of-the-art resources help our researchers and partners dive deep into biological processes to reveal new drug targets and promising therapies.
Programs
Our intensely collaborative environment brings together experts in complementary areas of biology to accelerate discoveries and clinical progress.
Research Themes
Four overarching themes focus our efforts on key questions and challenges in biomedical research.
Shared Resources
State-of-the-Art Instrumentation, Facilities and Services
Sanford Burnham Prebys has an extensive Shared Resources system with more than 20 core facilities. These facilities provide investigators with advanced technologies, expertise and instrumentation that may not be easily acquired or fully used by individual laboratories.
Our core facilities are staffed by expert technicians and provide high-quality interactive services for cost-effective sample analyses, as well as assistance in experiment design and grant or manuscript preparation. Many of these core services offer multiple levels of service assistance or hands-on investigator training for independent use of advanced instrumentation.
Let us help you
Most core facilities can provide services for outside researchers. Contact the facility manager or director for more information.