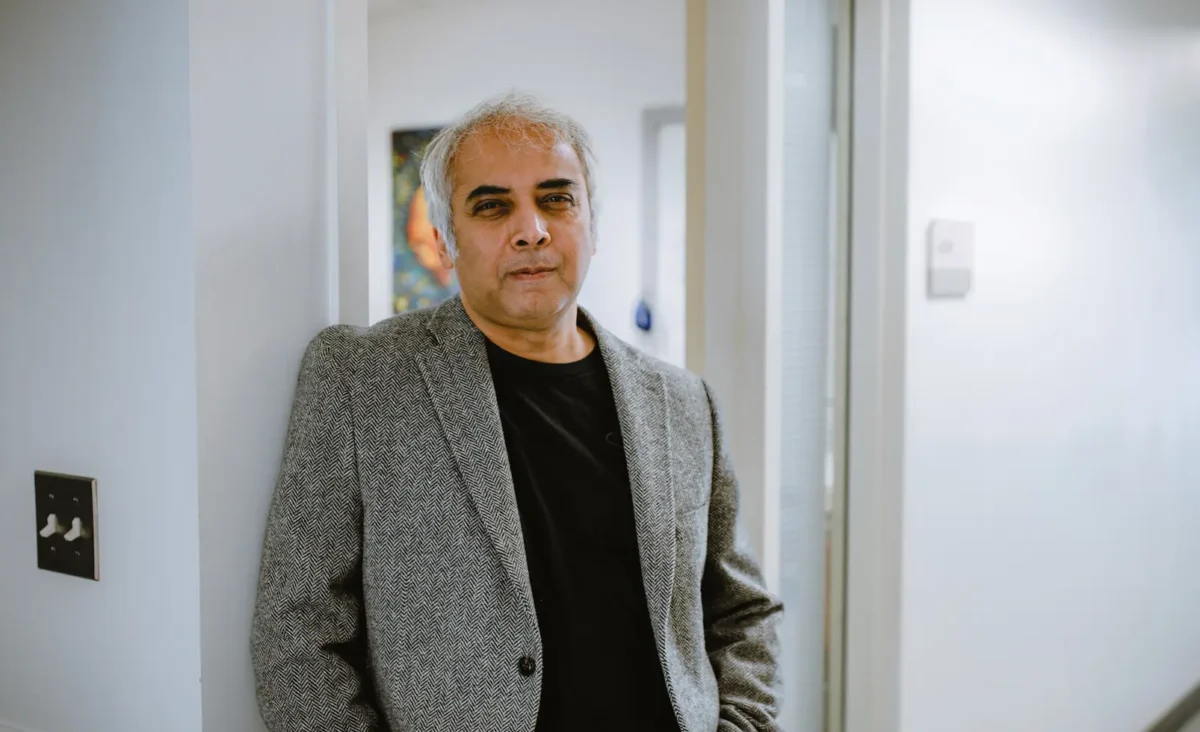
Dr. Bagchi comes to Sanford Burnham Prebys from the University of Minnesota, where he has been since 2008, having earlier held positions as assistant professor of Cell Biology & Development (Genetic Mechanism of Cancer) and Co-Director of Mouse Genetics Laboratory at University of Minnesota. Dr. Bagchi brings his lab with him and an established research program in genetic mechanisms of cancer. He completed his PhD degree at Jawaharlal Nehru University in New Delhi, India and then worked as a postdoctoral fellow at the Cold Spring Harbor Laboratory in New York. He has been an American Cancer Society Research Scholar since 2014 and Masonic Scholar since 2010.
Related Disease
Cancer
The Bagchi lab studies the genetic mutations that drive cancer. I am a geneticist by training, and I am passionate about finding the hidden drivers of cancer. In the research community, there has been tremendous progress in the last few decades in understanding the nature of the beast, but now is the time to translate those knowledge to actionable interventions. A riveting idea that has excited them for sometime is to illuminate the genetic mutations which act as life line to many cancers. This would expose their Achilles heel that we can exploit in near future to develop new forms of treatment. That is what the lab’s work is focused on. They have identified and characterized several key mutations that we have shown as proof of concept can be targeted in as much as 30% of all cancers. The focus of the lab is now to carry out experiments that will allow us to develop this work beyond the proof of concept, ideally to successful clinical trials.
 Sep 25, 2024
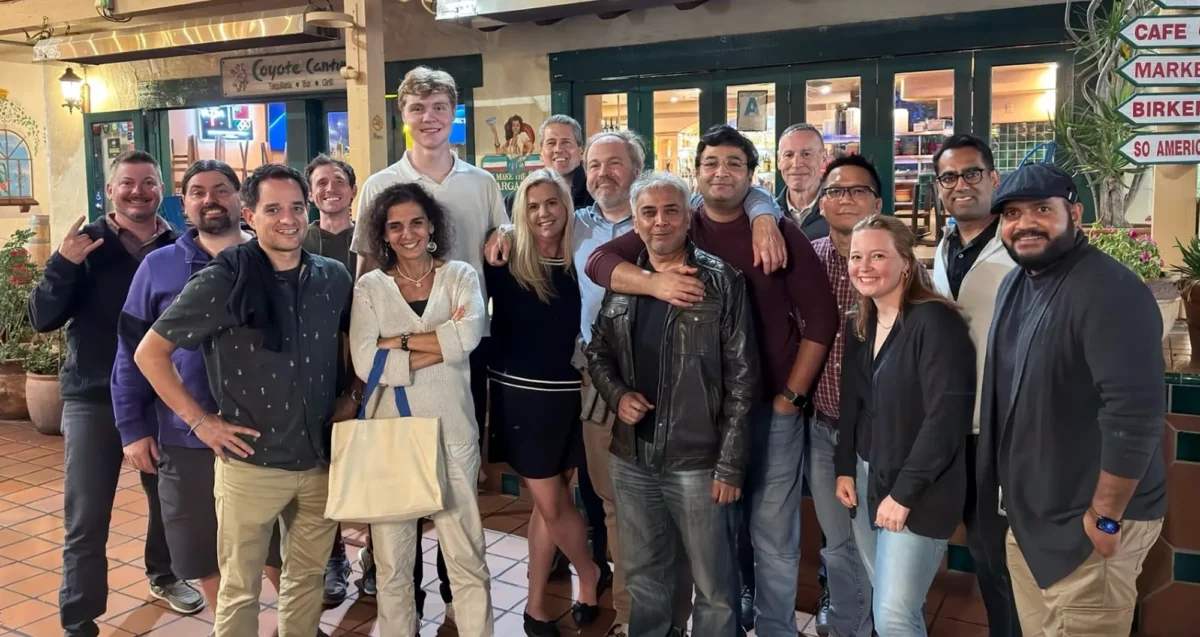
Sep 25, 2024Mammalian Genome Engineering Group holds 2024 symposium in San Diego
Sep 25, 2024Researchers convened at Sanford Burnham Prebys from September 12-15 to discuss research advances and new collaborations.
 Mar 20, 2023
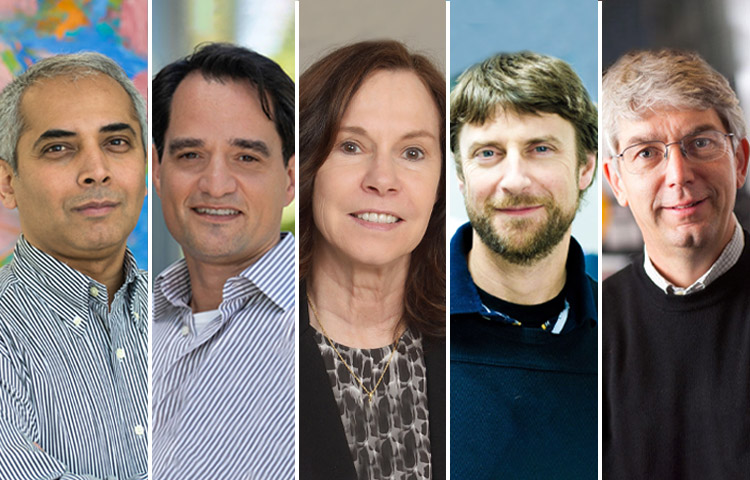
Mar 20, 2023Sanford Burnham Prebys researchers awarded Curebound grants
Mar 20, 2023Each year, Sanford Burnham Prebys joins Padres Pedal the Cause, an annual fundraising event that raises money for Curebound which…
 Aug 4, 2020
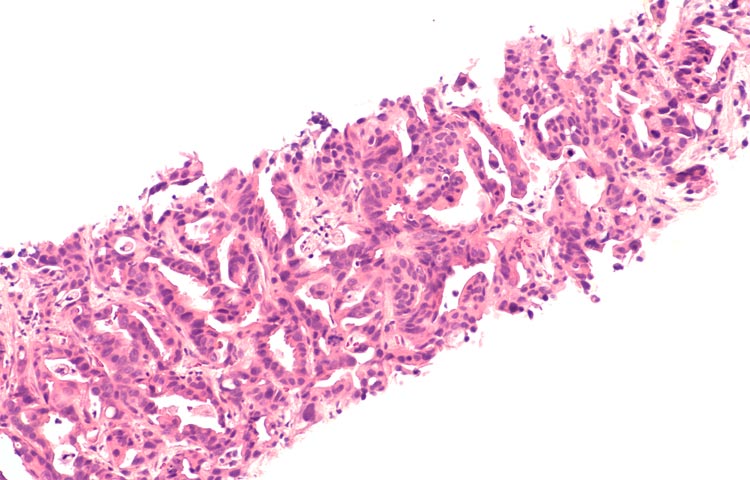
Aug 4, 2020Scientists discover novel drug target for pancreatic cancer
Aug 4, 2020Scientists have discovered a novel drug target that stops the deadly spread of pancreatic cancer Scientists at Sanford Burnham Prebys Medical Discovery Institute have
 Apr 6, 2017
Apr 6, 2017New investigator Anindya Bagchi joins Sanford Burnham Prebys Medical Discovery Institute
Apr 6, 2017
