Emerling received her B.A. from the University of California Santa Cruz and her PhD in molecular and cellular biology from Northwestern University. Emerling did her postdoctoral training at Harvard Medical School. She then became an Instructor of Cancer Biology in Medicine at Weill Cornell Medical College in New York City, where she continued her research on lipid kinase signaling and cancer metabolism. In August 2016, Brooke joined the faculty at Sanford Burnham Prebys Medical Discovery Institute as an Assistant Professor in the Cancer Metabolism and Signaling Networks Program.
Funding Awards and Collaborative Grants
Breast Cancer Research Foundation – AACR Career Development Award for Translational Breast Cancer Research
Mary Kay Foundation Innovative Translational Grant Award
Department of Defense Breast Research Program Breakthrough Award
Honors and Recognition
2014: NextGen Star – AACR Early-Career Speaker Award
2013-2016: Mastercard Ajay Banga Scientist Award
2013: AACR – Aflac Travel Fellowship Award
Related Disease
Breast Cancer, Cancer
Phenomena or Processes
Cancer Biology, Cancer Metabolism, Cell Signaling, Metabolic Processes, Signal Transduction
Research in the Emerling Lab is focused on understanding key signaling and metabolic pathways involved in the regulation of cellular function under pathological conditions such as cancer. Our research program centers around dissecting the roles of the family of non-canonical phosphatidylinositol kinases, called the phosphatidylinositol-5-phosphate 4-kinases (PI5P4Ks), in cancer metabolism using a multi-disciplinary approach integrating human, mouse, and worm models. Currently, a major research project in the Emerling Lab is determining the role of the PI5P4Ks in p53 mutant cancers, especially the triple-negative breast cancer subgroup where targeted therapies have not been effective.
 May 22, 2026
May 22, 2026Cholesterol-craving cancers need lipid enzymes to use metabolites for growth
May 22, 2026Study from the Emerling lab finds kinases move cholesterol in the cell to where it can activate a growth pathway…
 Mar 30, 2026
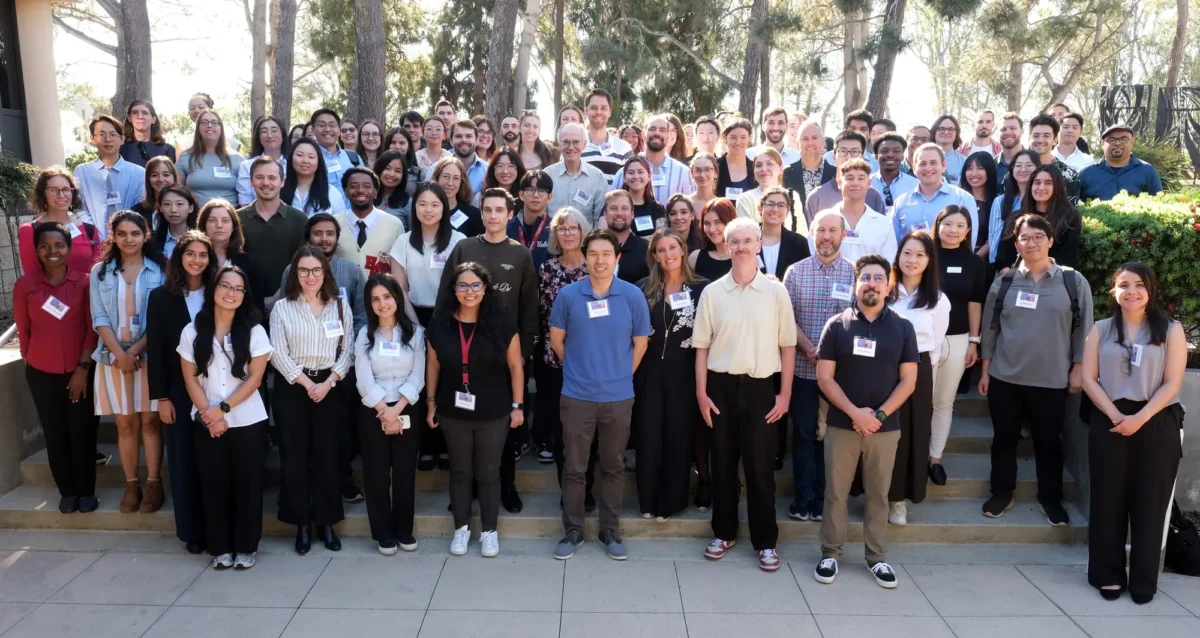
Mar 30, 2026Southern California metabolism scientists meet on the Mesa
Mar 30, 2026The fourth annual SoCal Metabolism Symposium brought together hundreds of experts and trainees to share the latest advances.
 Oct 21, 2025
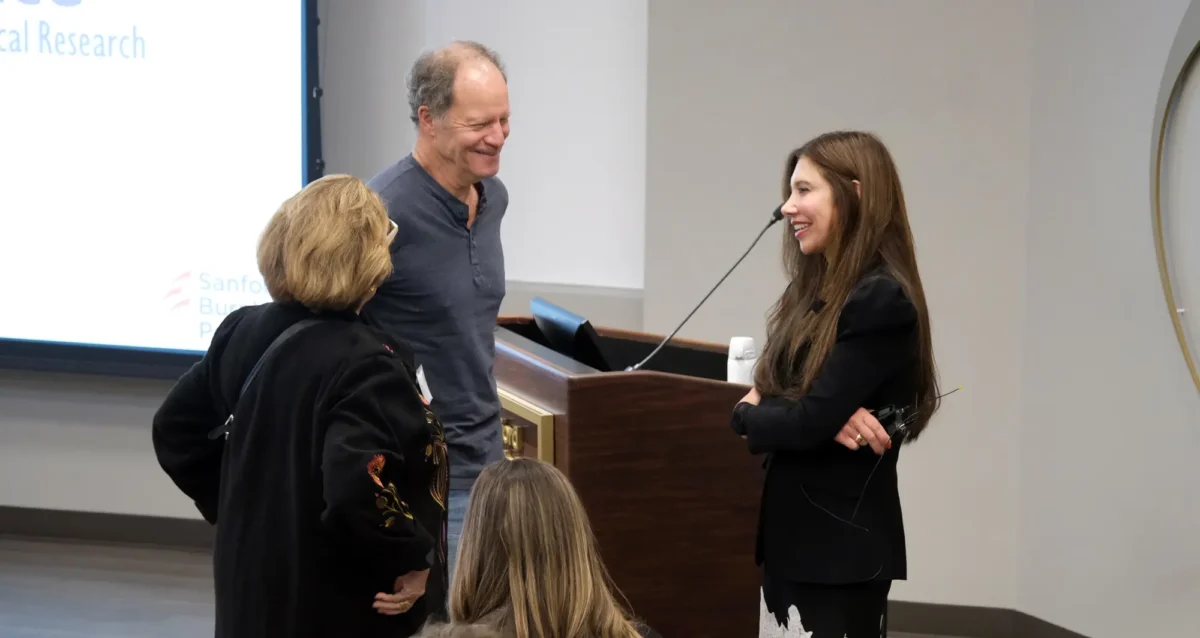
Oct 21, 2025Sanford Burnham Prebys hosts inaugural event in the Women in Science Lecture Series
Oct 21, 2025The series highlights the groundbreaking work and unique perspectives of women leaders in the biomedical sciences.
 Sep 11, 2025
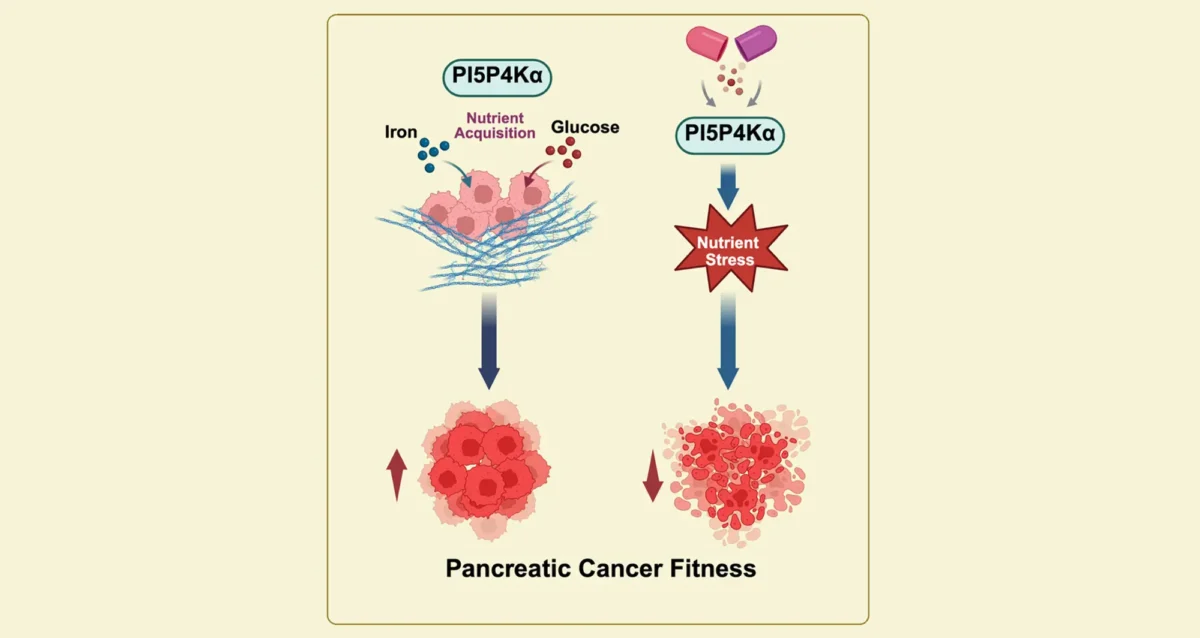
Sep 11, 2025Starving Cancer Cells of Essential Sugar and Iron
Sep 11, 2025Findings from Gurpreet Kaur Arora, Brooke Emerling, and Cosimo Commisso, identify key enzyme that pancreatic cancer cells need to obtain…
 Feb 12, 2025
Feb 12, 2025Curebound awards two grants to Sanford Burnham Prebys scientists
Feb 12, 2025The San Diego-based philanthropic organization has awarded $43 million in cancer research to date.
 Oct 18, 2024
Oct 18, 2024Two Sanford Burnham Prebys scientists selected for American Cancer Society postdoctoral fellowships
Oct 18, 2024The funds will support Alicia Llorente Lope and Ambroise Manceau who study breast and pancreatic cancer.

