Dr. Adams most recently led the Epigenetics Unit at the Beatson Institute for Cancer Research and the University of Glasgow, Institute of Cancer Sciences, in Scotland. He has also held positions at Wistar Institute (University of Pennsylvania), Drexel University and Fox Chase Cancer Center in Philadelphia.
Peter D. Adams obtained his BA in biochemistry at the University of Oxford, England and his PhD at Imperial Cancer Research Fund (now CR-UK). He did postdoctoral work with Dr. William G. Kaelin, Jr. at Dana-Farber Cancer Institute. Peter D. Adams is co-Editor-in-Chief of the journal Aging Cell.
Education
1993: PhD, Signal Transduction, Imperial Cancer Research Fund (CRUK), London, UK (Dr. Peter Parker, advisor)
1989: BA, Biochemistry, Oxford University, England
Honors and Recognition
2003-2008: Leukemia and Lymphoma Society Scholar
1999-2001: W.W. Smith Charitable Trust Fellowship
1999-2001: V Foundation Scholar
1995-1996: Cancer Research Foundation of America Fellowship
1993-1995: SERC/NATO Fellowship
1989: B.A. with Honors in Biochemistry, Oxford University, UK
1986: Awarded a Distinction in Oxford University Preliminary Examinations
1984-1989: 1984-1989: Exhibition holder for Academic Achievement at Oxford University, UK
1983: Lane Scholarship for Academic Achievement at King Henry VIII School, UK
Related Disease
Aging, Aging-Related Diseases, Colorectal Cancer, Leukemia/Lymphoma, Liver Cancer, Skin Cancer and Melanoma
Phenomena or Processes
Epigenetics
Dr. Adams’ lab investigates the impact of chromatin and epigenetics on cellular senescence, aging and cancer. In particular, the lab hypothesizes that age-associated changes in chromatin and epigenetic programming contribute to the dramatic age-associated increase in incidence of cancer. While age is the biggest single risk factor for most cancers, the reason for this is current poorly understood.
Dr. Adams’ research focus is driving epigenetics as a cross-cutting scientific theme and promoting access to single cell technologies at the Institute.
Peter D. Adams’ Research Report
We have mapped the epigenetic landscape of senescent cells, and are defining the causes and consequences of this altered landscape.
Cellular senescence is an irreversible proliferation arrest and pro-inflammatory phenotype triggered in primary cells by activated oncogenes and other molecular stresses. As a profound change in cell phenotype, the initiation and maintenance of senescence depends on reprogramming of chromatin, the epigenome and gene expression. Cellular senescence is a potent tumor suppressor mechanism. However, the accumulation of senescent cells with age also causes tissue aging, by blocking cell and tissue renewal and driving chronic inflammation. Indeed, in contrast to its acute tumor suppressive effects, chronic accumulation of inflammatory senescent cells is tumor promoting. In collaboration with Dr. Shelley Berger, we have mapped the distribution of several critical epigenetic regulators in proliferating and senescent cells, including DNA methylation, several histone modifications, histone variants and nuclear lamins. These collaborative studies have yielded critical insights into gene regulation in senescent cells (Cruickshanks et al., 2013; Rai et al., 2014; Shah et al., 2013), and the tumor suppressive and pro-aging effects of senescent cells (Cruickshanks et al., 2013; Nelson et al., 2016).
- Senescent cells harbour features of the cancer epigenome. Cruickshanks, H.A., McBryan, T., Nelson, D.M., Vanderkraats, N.D., Shah, P.P., van Tuyn, J., . . . Adams, P.D. Nat Cell Biol. 2013 Dec ;15(12):1495-506
- Mapping H4K20me3 onto the chromatin landscape of senescent cells indicates a function in control of cell senescence and tumor suppression. Nelson, D.M., Jaber-Hijazi, F., Cole, J.J., Robertson, N.A., Pawlikowski, J.S., Norris, K.T., Criscione, S.W., Pchelintsev, N.A., Piscitello, D., Stong, N., Rai, T.S., McBryan, T., ……Adams, P.D. Genome Biol. 2016 Jul 25;17(1):158
- HIRA orchestrates a dynamic chromatin landscape in senescence and is required for suppression of neoplasia. Rai, T.S., Cole, J.J., Nelson, D.M., Dikovskaya, D., Faller, W.J., Vizioli, M.G., . . . Adams, P.D. Genes Dev. 2014 Dec 15;28(24):2712-25
- Lamin B1 depletion in senescent cells triggers large-scale changes in gene expression and the chromatin landscape. Shah, P.P., Donahue, G., Otte, G.L., Capell, B.C., Nelson, D.M., Cao, K., . . . Adams*, P.D. Berger*, S.L. Genes Dev. 2013 Aug 15;27(16):1787-99. * Co-corresponding authors.
Landmark structure and functional studies on the HIRA histone chaperone complex and its role in senescence-mediated tumor suppression.
In collaboration with Dr. Ronen Marmorstein in Philadelphia we have dissected the structure-function relationships between HIRA and its binding partners, UBN1, CABIN1 and ASF1a and substrate histone H3.3 (Zhang et al., 2005). This included a crystal structure of the HIRA/ASF1a interaction surface and more recently the UBN1/histone H3.3 interaction surface (Tang et al., 2006). We were the first to describe the distribution of the HIRA complex across the mammalian epigenome (Pchelintsev et al., 2013). In functional studies, we have demonstrated the role of this DNA replication independent histone chaperone complex in the control of chromatin in non-proliferating senescent cells (Rai et al., 2014). These studies have been facilitated by the mouse monoclonal and rabbit polyclonal antibodies that we have made to all subunits of the complex. More recently, we have generated the first conditional knock out mice of HIRA, UBN1 and CABIN1 and are using these to establish in vivo functions (Rai et al., 2014). Of particular note, we have revealed a function for HIRA in promoting healthy aging and the suppression of cancer (Rai et al., 2014).
- Formation of MacroH2A-containing senescence-associated heterochromatin foci and senescence driven by ASF1a and HIRA. Zhang, R., Poustovoitov, M.V., Ye, X., Santos, H.A., Chen, W., Daganzo, S.M., . . . Adams, P.D. Dev Cell. 2005 Jan ;8(1):19-30
- Structure of a human ASF1a-HIRA complex and insights into specificity of histone chaperone complex assembly. Tang, Y., Poustovoitov, M.V., Zhao, K., Garfinkel, M., Canutescu, A., Dunbrack, R., . . . Adams*, P.D., Marmorstein*, R. (2006) Nat Struct Mol Biol. 2006 Oct ;13(10):921-9
- Placing the HIRA histone chaperone complex in the chromatin landscape. Pchelintsev, N.A., McBryan, T., Rai, T.S., van Tuyn, J., Ray-Gallet, D., Almouzni, G., and Adams, P.D. (2013). Cell Rep. 2013 Apr 25;3(4):1012-9
- HIRA orchestrates a dynamic chromatin landscape in senescence and is required for suppression of neoplasia. Rai, T.S., Cole, J.J., Nelson, D.M., Dikovskaya, D., Faller, W.J., Vizioli, M.G., . . . Adams, P.D. (2014). Genes Dev. 2014 Dec 15;28(24):2712-25
We coined the term “chromostasis” to describe the presumptive homeostatic mechanisms that confer epigenetic stability over the lifecourse, and we were major contributors to the first demonstration of a DNA methylation clock in the mouse.
Maintenance of cell phenotype and suppression of disease, including cancer, over the lifecourse depends on a high level of epigenetic stability. However, since chromatin is inherently dynamic (Rai et al., 2014), this steady state stability likely reflects a challenge for the cell. Therefore, presumptive “chromatin homeostasis” or “chromostasis” mechanisms are predicted to actively maintain an epigenetic steady state over the lifecourse, thereby suppressing age-associated disease (Rai et al., 2014). We have shown that histone chaperone HIRA is one such factor that contributes to epigenetic stability in non-proliferating cells (Ye et al., 2007). Recently, we reported the first DNA methylation clock in the mouse, and showed that diverse interventions – genetic, dietary and drug – that promote longevity of mice also suppress age-associated epigenetic changes and slow progression of this DNA methylation “clock”, i.e. enhance chromostasis (Cole et al., 2017; Wang et al., 2017).
- HIRA orchestrates a dynamic chromatin landscape in senescence and is required for suppression of neoplasia. Rai, T.S., Cole, J.J., Nelson, D.M., Dikovskaya, D., Faller, W.J., Vizioli, M.G., . . . Adams, P.D. (2014). Genes Dev. 2014 Dec 15;28(24):2712-25
- Formation of MacroH2A-containing senescence-associated heterochromatin foci and senescence driven by ASF1a and HIRA. Zhang, R., Poustovoitov, M.V., Ye, X., Santos, H.A., Chen, W., Daganzo, S.M., . . . Adams, P.D. Dev Cell. 2005 Jan ;8(1):19-30
- Diverse interventions that extend mouse lifespan suppress shared age-associated epigenetic changes at critical gene regulatory regions. Cole, J.J., Robertson, N.A., Rather, M.I., Thomson, J.P., McBryan, T. Sproul, D. Wang, T. Brock, C; Clark, W., Ideker, T. Meehan, R.R. Miller, R.A., Brown-Borg, H., Adams, P.D. Genome Biol. 2017 Mar 28;18(1):58
- Epigenetic aging signatures in mice are slowed by dwarfism, calorie restriction and rapamycin treatment. Wang, T., Tsui, B., Kreisberg, J.F., Robertson, N.A., Gross, A.M., Carter, H., Brown-Borg, H., Adams, P.D, Ideker, T. Genome Biol. 2017 Mar 28;18(1):57
We have demonstrated a “tug of war” between tumor suppressive oncogene-induced senescence and oncogenic activated Wnt signaling in melanocytic neoplasia (Adams and Enders, 2008; Ye et al., 2007).
The balance between these tumor suppressive and oncogenic activities determines the efficiency of senescence-mediated tumor suppression. For example, we showed that in oncogene-expressing melanocytes a low level of activated Wnt signaling promotes benign nevus formation (Pawlikowski et al., 2013). However, a high level of activated Wnt signaling, caused by germline sequence variants, promotes giant congenital nevi in the form of congenital melanocytic nevus (CMN) syndrome (Pawlikowski et al., 2015). In a mouse model that closely recapitulates the human genetics, we showed that activated Wnt signaling and an activated Ras oncogene (NRasQ61K) cooperate to drive CMN syndrome, and that this is suppressed by acute post-natal treatment with MEK inhibitors (Pawlikowski et al., 2015). Based on these studies, our collaborator Dr. Veronica Kinsler is preparing to test MEK inhibitors in babies afflicted by CMN syndrome.
- Downregulation of Wnt signaling is a trigger for formation of facultative heterochromatin and onset of cell senescence in primary human cells. Ye, X., Zerlanko, B., Kennedy, A., Banumathy, G., Zhang, R., and Adams, P.D. Mol Cell. 2007 Jul 20;27(2):183-196
- Acute Inhibition of MEK Suppresses Congenital Melanocytic Nevus Syndrome in a Murine Model Driven by Activated NRAS and Wnt Signaling. Pawlikowski, J.S., Brock, C., Chen, S.C., Al-Olabi, L., Nixon, C., McGregor, F., . . . Adams, P.D. J Invest Dermatol. 2015 Aug;135(8):2093-2101
- Wnt signaling potentiates nevogenesis. Pawlikowski, J.S., McBryan, T., van Tuyn, J., Drotar, M.E., Hewitt, R.N., Maier, A.B., . . . Adams, P.D. Proc Natl Acad Sci U S A. 2013 Oct 1;110(40):16009-14
- Wnt signaling and senescence: A tug of war in early neoplasia? Adams, P.D., and Enders, G.H. Cancer Biol Ther. 2008 Nov;7(11):1706-11
We first characterized Cytoplasmic Chromatin Fragments (CCF) in senescent cells and in collaboration defined a role for CCF as drivers of inflammation via the cGAS/STING cytoplasmic DNA sensing anti- viral pathway.
Cellular senescence is a potent tumor suppressor mechanism by virtue of proliferation arrest and the senescence associated secretory phenotype (SASP) which promotes clearance of pre-malignant cells by the immune system. However, the mechanism responsible for initiation of SASP has been unknown. In 2013, our lab first characterized and named CCF as fragments of chromatin expelled from the nucleus of senescent cells into the cytoplasm (Ivanov et al., 2013). Then, in 2015, in collaboration with Shelley Berger’s laboratory, we showed that formation of CCF depends on interaction of lamin B1 and autophagy adaptor LC3 in the nucleus, and that lamin B1 is a nuclear substrate of autophagy (Dou et al., 2015). Most recently, again with Shelley Berger’s lab, we have shown that CCF are sensed by the cytoplasmic DNA sensing anti-viral apparatus, cGAS and STING, and this leads to activation of NFkB and SASP in senescent cells (Dou et al., 2017). The role of CCF as triggers of SASP, via cGAS and STING, has been confirmed by several other labs. Most recently, we showed that formation of CCF is triggered by a retrograde mitochondria-to-nucleus signalling pathway that constututes a target for candidate anti-SASP healthy aging interventions (Vizioli et al., 2020).
- Lysosome-dependent processing of histones in senescent cells. Ivanov, A., Pawlikowski, J., Manoharan, I., van Tuyn, J., Nelson, D. M., Rai, T. S., Shah, P. P., Drotar, M., Wu, H., Berger, S. L., and Adams, P.D. J. Cell Biol. 2013 July 1; 202 (1): 129–143
- Autophagy mediates degradation of nuclear lamina. Dou, Z., Capell, BC., Drake, AM., Shah, PP., Dorsey, J., Simola, D., Donahue, G., Zhu, Z., Sammons, M., Rai, TS., Natale, C., Ridky, T., Goldman, R., Adams, P.D., Berger, SL. Nature. 2015 Oct 28; 527:105-109
- Cytoplasmic chromatin triggers inflammation in senescence and cancer. *Corresponding authors. Dou, D., Ghosh, K., Vizioli, MG., Zhu, J., Sen, P., Wangensteen, KJ., Simithy, J., Lan, Y., Lin, Y., Zhou, Z., Capell, BC., Xu, C., Xu, M., Kieckhaefer, JE., Jiang, T., Shoshkes-Carmel, M., Ahasan Al Tanim, KM., Barber, GN., Seykora, JT., Millar, SE., Kaestner, KH., Garcia, BA., Adams, P.D.*, Berger, SL*. Nature. 2017 Oct 4; 550:402-406
- Mitochondria-to-nucleus retrograde signaling drives formation of cytoplasmic chromatin and inflammation in senescence. Vizioli, M.G., Liu, T., Miller, K.N., Robertson, N., Gilroy, K., Lagnado, A., Perez-Garcia, A., Kiourtis, C., Dasgupta, N., Lei, X., Kruger, P.J., Nixon, C., Clark, W., Jurk, D., Bird, T.G., Passos, J.F., Berger, S.L., Dou, Z., Adams, P.D. Genes and Dev. 2020 Jan 30; 34, 428-445
 Jun 8, 2026
Jun 8, 2026CIAO Study: 11th annual longevity symposium provides new peeks and possibilities at longer, healthier lives
Jun 8, 2026The Cilento Initiative on Aging Outcomes (CIAO) held its 11th annual research symposium at Sanford Burnham Prebys, with more than a…
 Jun 3, 2026
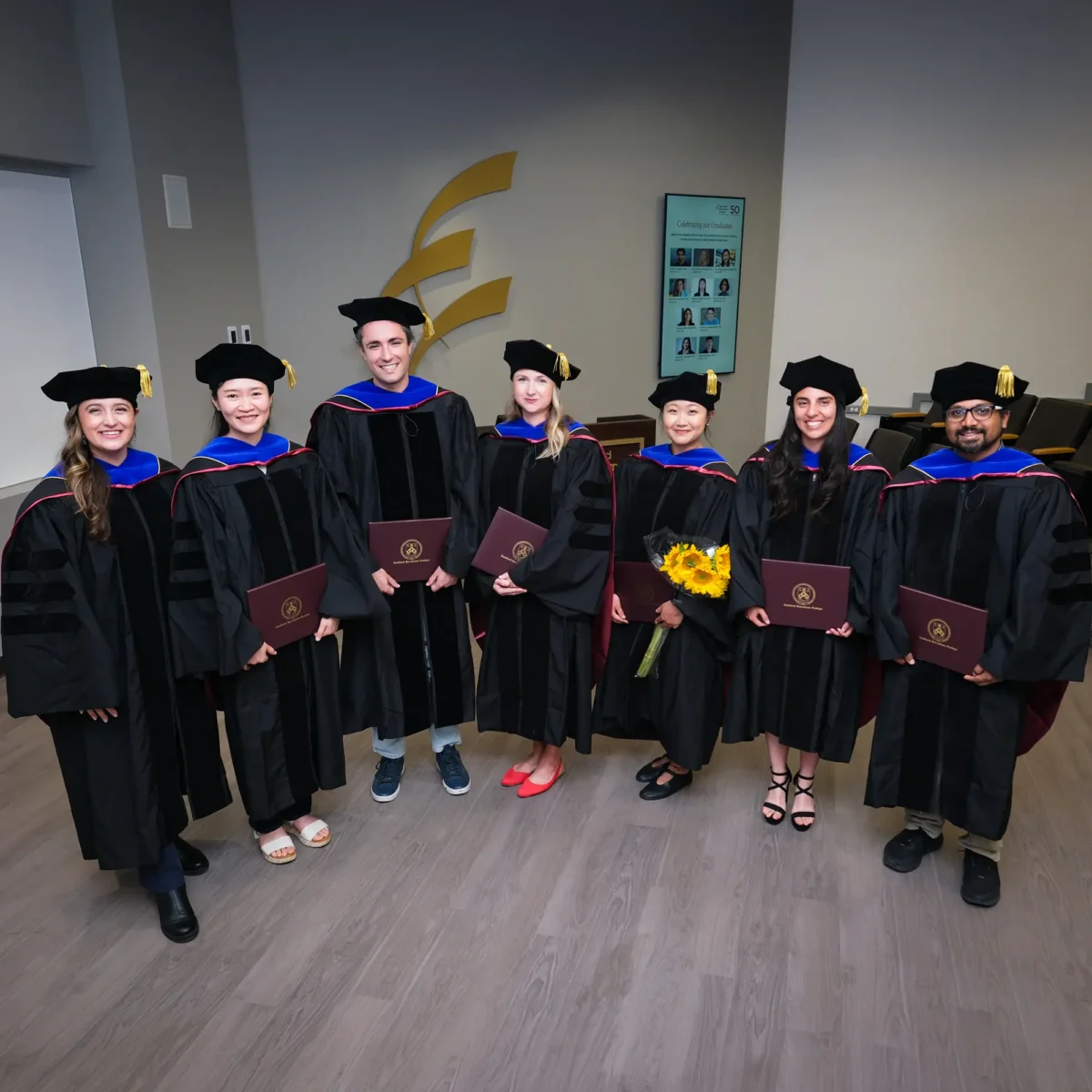
Jun 3, 2026Sanford Burnham Prebys recognizes 10 doctoral degree recipients
Jun 3, 2026The Institute’s Graduate School of Biomedical Sciences held its third Commencement ceremony to celebrate new alumni.
 Mar 25, 2026
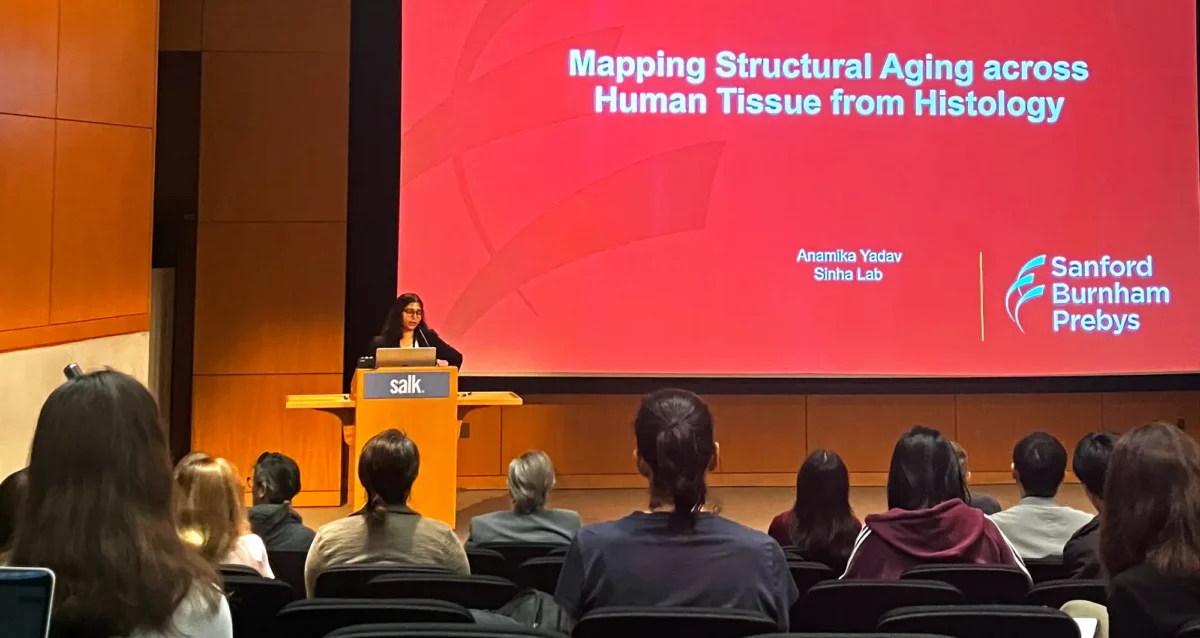
Mar 25, 2026La Jolla research meeting convenes scientists studying aging
Mar 25, 2026Scientists gathered at the La Jolla Aging Meeting on March 19, 2026, in the Salk Institute’s Conrad T. Prebys Auditorium.
 Oct 6, 2025
Oct 6, 2025Sanford Burnham Prebys expert surveys science on how to treat the most common brain cancer
Oct 6, 2025New editorial recommends a multimodal perspective examining glioblastoma from tumor biology through to surgery.
 Apr 1, 2025
Apr 1, 2025Experts exchange advances in the science of healthier aging in San Diego
Apr 1, 2025Two scientific meetings in late March brought together researchers studying aging and its implications for disease.
 Mar 14, 2025
Mar 14, 2025Cellular circuit controls how DNA damage is repaired, affecting risk of disease as we age
Mar 14, 2025Study reveals new information about how to prevent chronic inflammation from zombie-like cells that accumulate with age.
