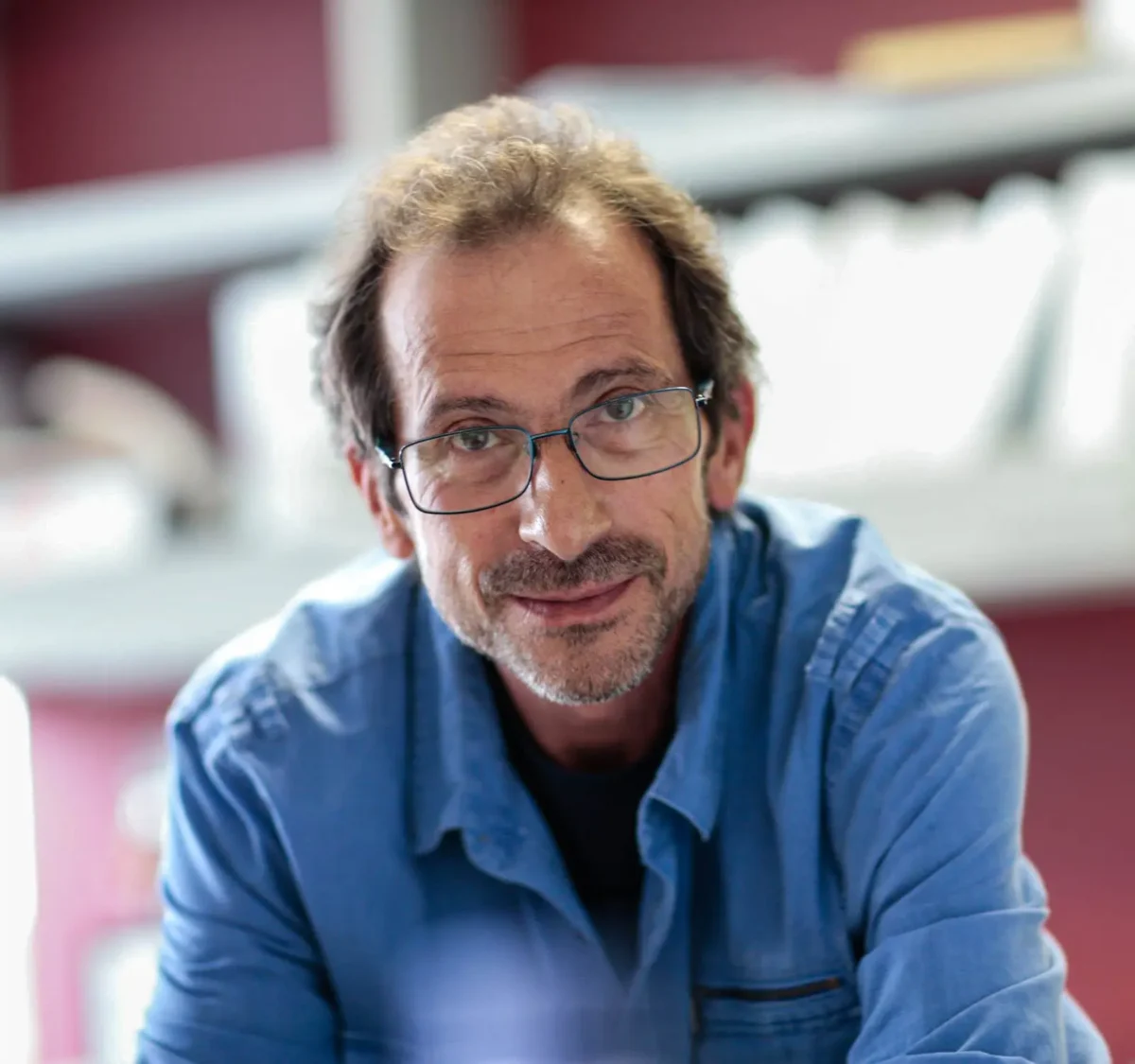
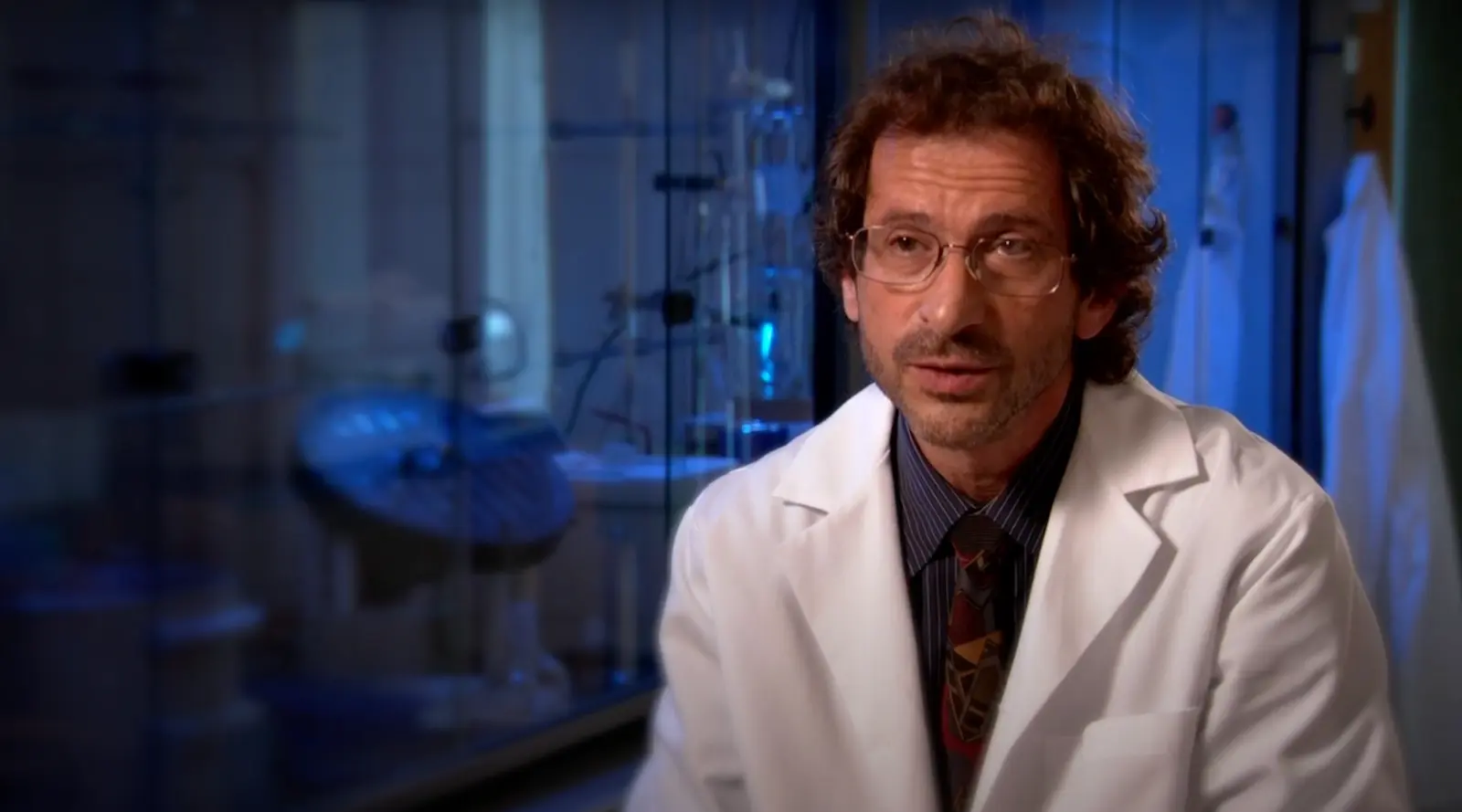
Andrei Osterman is a Professor in the Immunity and Pathogenesis Program Program at the Infectious and Inflammatory Disease Center of Sanford Burnham Prebys (since August 2003). He received his doctorate from Moscow State University in 1983, did postdoctoral work UT Southwestern Medical Center, and held the position of the Director and then Vice President of Research at Integrated Genomics in 1999-2003. Dr. Osterman is one of the founders of the Fellowship for Interpretation of Genomes (FIG), a nonprofit research organization that launched the Project to Annotate 1,000 Genomes in 2003. FIG provides the open-source integration of all publicly available genomes and tools for their comparative analysis, annotation, and metabolic reconstruction.
Related Disease
Breast Cancer, Cancer, Infectious Diseases, Radiation Damage, Skin Cancer and Melanoma
The main focus of Dr. Osterman’s research team is on fundamental and applied aspects of the key metabolic subsystems in a variety of species, from bacteria to human. This group uses a systems biology approach to reconstruct and explore metabolic and transcriptional regulatory networks. This approach combines comparative genomics and other bioinformatic techniques with biochemical and genetic experiments for pathway, gene and target discovery. Using this approach this group predicted and experimentally verified numerous enzyme families in the metabolism of cofactors, carbohydrates, and amino acids. Recent breakthroughs included prediction and characterization of novel transporters, transcriptional regulators and carbohydrate utilization pathways in a number of model bacterial systems. Applications in the field of infectious disease include identification of novel drug targets and structure-based development of novel anti-infective agents. New directions in cancer research are based on application of metabolic profiling technology for identification of novel diagnostic and therapeutic targets. Other directions of the on-going research include bioinformatics of regulatory proteolysis and applications of structural modeling for exploration of metabolic networks and gene discovery.
 Jun 3, 2026
Jun 3, 2026Sanford Burnham Prebys recognizes 10 doctoral degree recipients
Jun 3, 2026The Institute’s Graduate School of Biomedical Sciences held its third Commencement ceremony to celebrate new alumni.
 Nov 7, 2025
Nov 7, 2025Evolving antibiotic resistance under pressure
Nov 7, 2025Study shows how a bacterial pathogen avoids biting the dust through genomic mutations, may lead to personalized treatment.
 Jul 16, 2025
Jul 16, 2025Bacterial genomes hold clues for creating personalized probiotics
Jul 16, 2025Studying the genomes of beneficial bacteria may lead to new targeted probiotic treatments.
 Oct 2, 2024
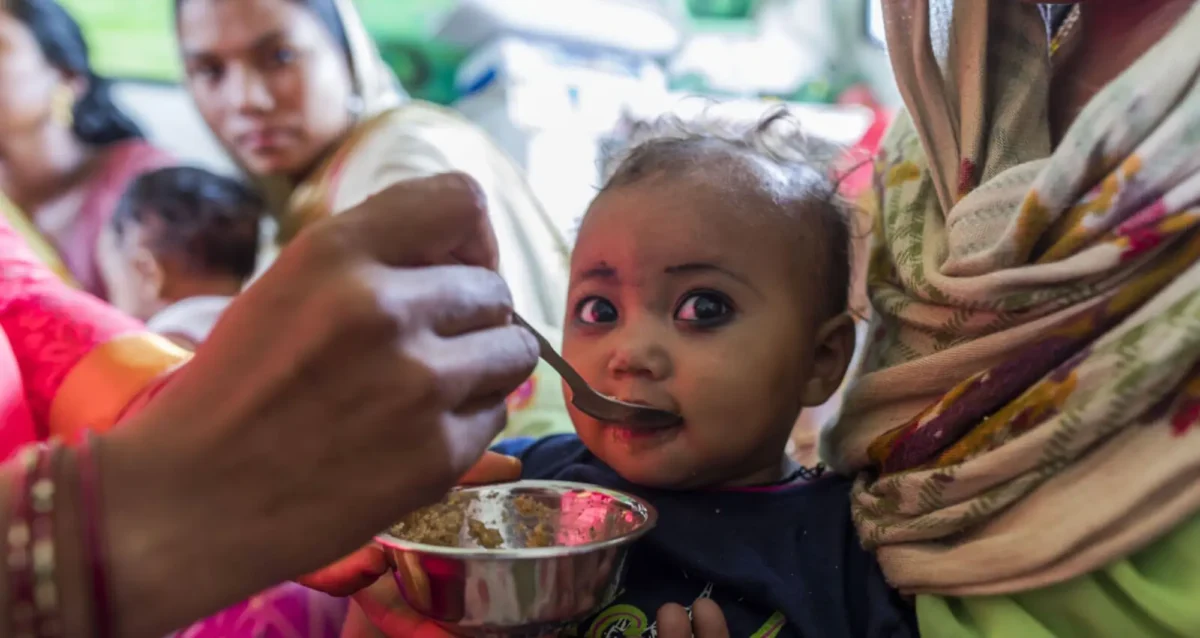
Oct 2, 2024Gut microbiome repair in children with severe acute malnutrition
Oct 2, 2024Researchers around the world, including Andrei Osterman, have been investigating potential remedies for child malnutrition.
 Aug 13, 2024
Aug 13, 2024Dodging AI and other computational biology dangers
Aug 13, 2024Sanford Burnham Prebys scientists say that understanding the potential pitfalls of using artificial intelligence and computational biology techniques in biomedical…
 Aug 8, 2024
Aug 8, 2024Scripting their own futures
Aug 8, 2024At Sanford Burnham Prebys Graduate School of Biomedical Sciences, students embrace computational methods to enhance their research careers

